Network Generator
ndtools.network_generator creates synthetic network datasets (nodes, edges, probabilities)
that conform to this repo’s schemas.
Quick Start (Windows cmd)
You can call the generator from ndtools:
from pathlib import Path
from ndtools.network_generator import GenConfig, generate_and_save
cfg = GenConfig(
name="ws_n60_k6_b015",
generator="ws",
description="WS n=60 k=6 beta=0.15",
generator_params={"n_nodes": 60, "k": 6, "p_ws": 0.15, "p_fail": 0.1},
seed=7,
)
repo_root = Path(__file__).resolve().parents[1]
out_base = repo_root / "generated"
ds_root = generate_and_save(out_base, cfg, draw_graph=True)
print("Wrote:", ds_root)
Available Network Types
Overview
Network type |
Typical real-world interpretation |
Generative principle |
Key structural feature |
|---|---|---|---|
Grid |
Planned grid-layout road networks |
Regular lattice |
Uniform degree |
Erdős–Rényi (ER) |
Purely random connectivity |
Random edge formation |
Homogeneous degree distribution |
Watts–Strogatz (WS) |
Small-world networks (e.g. social or collaboration networks) |
Local lattice with random rewiring |
High clustering, short paths |
Random geometric |
Physical infrastructure networks constrained by distance (e.g. roads, pipelines, power distribution) |
Spatial proximity |
Spatial clustering |
Configuration graph |
Networks preserving observed node connectivities while randomising links |
Fixed number of connections per node |
Non-uniform number of connections across nodes |
Barabási–Albert (BA) |
Hub-based networks formed by growth (e.g. airline, internet, service networks) |
Growth with preferential attachment |
Scale-free degree distribution with hubs |
Example networks are illustrated below:
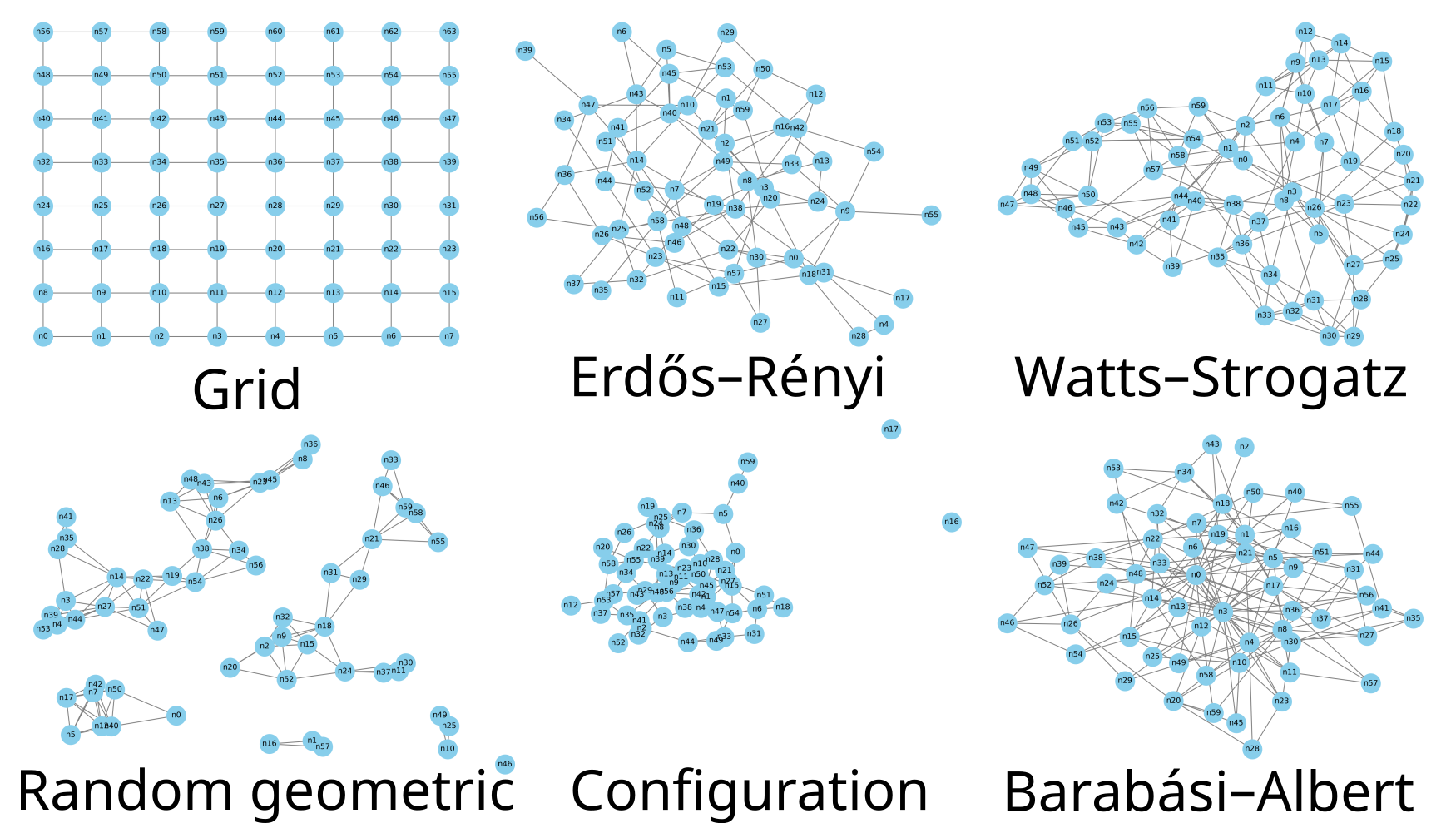
Figure 2. Example networks generated by the random network generator.
- where the models are generated with parameters:
Grid (8×8),
Erdős–Rényi (n=60, p=0.05),
Watts–Strogatz (n=60, k=6, β=0.15),
Random Geometric (n=60, r=0.17),
Configuration model (n=60, avg_deg=3),
Barabási–Albert (n=60, m=3).
Notes on Edge Counts
ER: expected edges \(E \approx p \cdot \frac{n(n-1)}{2}\).
WS: edges fixed by
k: \(E = \frac{n k}{2}\) (β changes structure, not count).BA: edges fixed by
m: \(E = m n - \frac{m(m+1)}{2}\).RG: edges grow roughly with \(r^2\); tune
--radius(e.g.,0.17for ~150 edges atn=60).Config: edges follow the synthesized degree sequence;
avg_deg≈2E/n.
Network generator parameters
The high-level entry point is generate_and_save(out_base, config, ...),
which (i) generates a network, (ii) assigns edge failure probabilities,
(iii) saves the dataset, (iv) optionally validates schema, visualises the graph,
and updates the registry.
Common (applies to all generators)
These parameters are read from config and affect all network types.
out_base(Path)Output base directory. The dataset is saved under this path.
config.generator(str)Generator type. Supported values:
grid|lattice|erdos_renyi|er|watts_strogatz|ws|barabasi_albert|ba|configuration|config|random_geometric|rg.config.name(str)Dataset name (used for the dataset folder).
config.version(str)Dataset version label used in the output path.
config.description(str)Human-readable description written to metadata/README.
config.seed(int)Random seed for reproducibility (used by all generators except
grid/lattice).config.generator_params(dict)Dictionary of generator parameters (see “Generator-specific parameters” below).
p_fail(float; insideconfig.generator_params)Edge failure probability used by
assign_edge_probs. Survival probability is1 - p_fail. If omitted, defaults to0.1.
Generator-specific parameters
These live inside config.generator_params and depend on config.generator.
Grid / Lattice (grid or lattice)
rows(int)Number of grid rows.
cols(int)Number of grid columns.
Example usage:
from pathlib import Path
from ndtools.network_generator import GenConfig, generate_and_save
cfg = ng.GenConfig(
name="grid_4x3",
generator="grid",
description="test grid",
generator_params={"rows": 4, "cols": 3, "p_fail": 0.1},
seed=None, # grid is deterministic
)
repo_root = Path(__file__).resolve().parents[1]
out_base = repo_root / "generated"
ds_root = generate_and_save(out_base, cfg, draw_graph=True)
print("Wrote:", ds_root)
Erdős–Rényi (erdos_renyi or er)
n_nodes(int)Number of nodes.
p(float)Edge probability.
Example usage:
from pathlib import Path
from ndtools.network_generator import GenConfig, generate_and_save
cfg = ng.GenConfig(
name="er_n20_p03",
generator="er",
description="test er",
generator_params={"n_nodes": 20, "p": 0.3, "p_fail": 0.1},
seed=7,
)
repo_root = Path(__file__).resolve().parents[1]
out_base = repo_root / "generated"
ds_root = generate_and_save(out_base, cfg, draw_graph=True)
print("Wrote:", ds_root)
Watts–Strogatz (watts_strogatz or ws)
n_nodes(int)Number of nodes.
k(int)Each node is connected to
knearest neighbours in ring topology (typically even).p_ws(float)Rewiring probability (β). Internally mapped to
p_rewirefor the underlying generator function.
Example usage:
from pathlib import Path
from ndtools.network_generator import GenConfig, generate_and_save
cfg = ng.GenConfig(
name="ws_n60_k6_b015",
generator="ws",
description="test ws",
generator_params={"n_nodes": 60, "k": 6, "p_ws": 0.15, "p_fail": 0.1},
seed=7,
)
repo_root = Path(__file__).resolve().parents[1]
out_base = repo_root / "generated"
ds_root = generate_and_save(out_base, cfg, draw_graph=True)
print("Wrote:", ds_root)
Barabási–Albert (barabasi_albert or ba)
n_nodes(int)Number of nodes.
m(int)Number of edges to attach from each new node to existing nodes.
Example usage:
from pathlib import Path
from ndtools.network_generator import GenConfig, generate_and_save
cfg = ng.GenConfig(
name="ba_n60_m3",
generator="ba",
description="test ba",
generator_params={"n_nodes": 60, "m": 3, "p_fail": 0.1},
seed=7,
)
repo_root = Path(__file__).resolve().parents[1]
out_base = repo_root / "generated"
ds_root = generate_and_save(out_base, cfg, draw_graph=True)
print("Wrote:", ds_root)
Configuration model (configuration or config)
n_nodes(int)Number of nodes.
avg_deg(float)Target average degree.
Example usage:
from pathlib import Path
from ndtools.network_generator import GenConfig, generate_and_save
cfg = ng.GenConfig(
name="config_n60_deg3p2",
generator="config",
description="test config",
generator_params={"n_nodes": 60, "avg_deg": 3.2, "p_fail": 0.1},
seed=7,
)
repo_root = Path(__file__).resolve().parents[1]
out_base = repo_root / "generated"
ds_root = generate_and_save(out_base, cfg, draw_graph=True)
print("Wrote:", ds_root)
Random Geometric (random_geometric or rg)
n_nodes(int)Number of nodes.
radius(float)Connection radius (typically in
[0, 1]depending on your implementation).
Example usage:
from pathlib import Path
from ndtools.network_generator import GenConfig, generate_and_save
cfg = ng.GenConfig(
name="rg_n60_r017",
generator="rg",
description="test rg",
generator_params={"n_nodes": 60, "radius": 0.17, "p_fail": 0.1},
seed=7,
)
repo_root = Path(__file__).resolve().parents[1]
out_base = repo_root / "generated"
ds_root = generate_and_save(out_base, cfg, draw_graph=True)
print("Wrote:", ds_root)
Pipeline options (not network-specific)
These are keyword arguments to generate_and_save that affect saving/validation/visualisation,
not the graph topology itself.
update_registry_flag(bool, defaultFalse)If
True, updateregistry.jsonat the repository root with the new dataset entry.
Visualisation options
draw_graph(bool, defaultTrue)If
True, attempt to render a graph image from saved data.graph_layout(str, default"spring")Layout name passed to the drawing function.
graph_name(str, default"graph.png")Output image filename.
graph_kwargs(dict or None)Extra keyword arguments forwarded to the drawing function.
Schema validation
schema_dir(Path or None)If provided, validate the generated dataset against the schema in this directory.
Generated Datasets
Data Structure
Example generated datasets are available under:
generated/<name>/v1/
data/
nodes.json
edges.json
probs.json
graph.png (if plotting enabled)
README.md
metadata.json
File Details
nodes.json(dict){"n0": {"x": <float|null>, "y": <float|null>}, ...}Grid assigns integer lattice coordinates (
x = i % cols,y = i // cols).ER / WS / BA / Config set
x,ytonull(no embedded coordinates).RG sets positions from the unit-square coordinates used to build the graph.
edges.json(dict){"e0": {"from": "n0", "to": "n1", "directed": false}, ...}probs.json(dict)Binary edge state probabilities (failure/survival) per edge id:
{ "e0": {"0": {"p": 0.1}, "1": {"p": 0.9}}, ... }
graph.png(optional)A preview figure rendered by
ndtools.graphs.draw_graph_from_data(). If nodes have numericx,y(e.g., RG, Grid), those are used; otherwise a layout is computed.
Acknowledgments
The network generator extensions were drafted by Alex Sixie Cao.